Software dedicated to protein scientists
Use our modules for proteins to register and align sequences, select best antibodies, track protein expression, purification and characterization, and share data with colleagues in real time.
Our digital solutions facilitate protein management: from antibody selection to protein production, all information is gathered in a single database to ease your daily life.

Register proteins
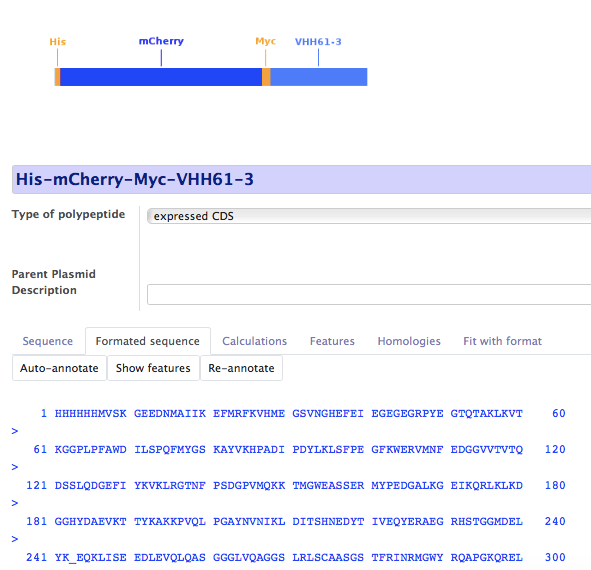
Yu-Protein is a Biologics Registration system. It has advanced features for protein format description and analysis. They are especially useful for antibody sequence analysis, but apply to any Biologics, without limitation.
Automatic protein translation and annotation
New proteins are compared to existing ones to ensure uniqueness and a new Compound Number is attributed.
Analysis of protein sequence is made according to the declared format. In the case of an antibody, the closest germline sequence is found and CDRs are identified based on the alignment.
Taking advantage of the relational database, coding sequences are automatically deduced from expression vectors.
Track protein expression, purification and characterization
Specify your expression system: bacterial, mammalian cell lines, yeast, hybridoma or recombinant hosts.
Indicate expression results (concentration of produced protein, volume, etc).
Enter purification steps, buffer used, final concentration of protein batches, tag cleavage.
Share data within a group of scientists through a single database.
Create aliquots and store them in various locations.
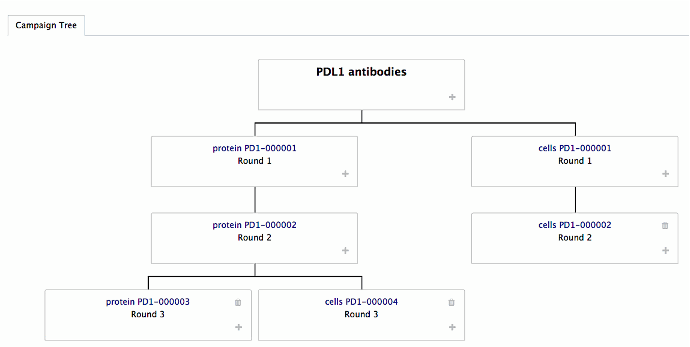
Antibody screening module
Build your campaign and selection strategy as a tree ( see picture)
Create a new selection arm direclty from the tree
Enrichment factors are automatically calculated
ELISA results: import absorbance in 96-well or 384 well-plate format from readers and store results in database.
Create ELISA templates (choose specific place for controls)
Visual display of positive clones
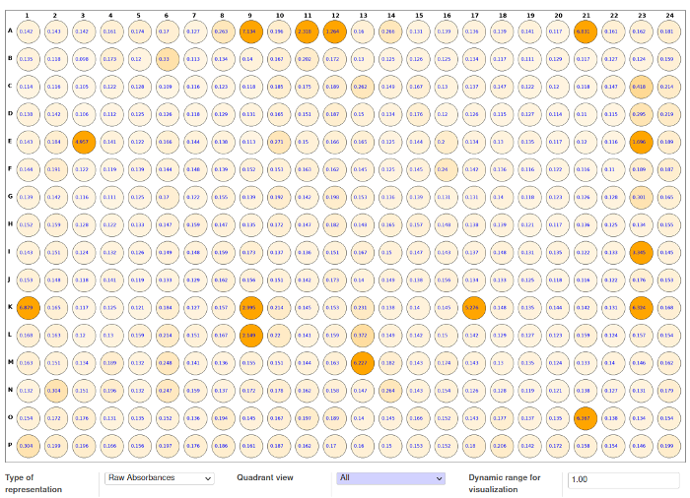
Creation of Hit-List: data-driven decision
Monitor antibody selection campaign thanks to Yu-Select Overview, an automatically generated table gathering all information
Filters can be positioned to create a selection of antibodies of interest and store it as 'Hit-List'
Create Hit-Lists at different stages of antibody selection process
Re-array a Hit-List to secure the best clones in a new glycerol plate (before ELISA confirmation, re-sequencing, reformatting...)
Reformat a 'Hit-List': antibody sequences are pasted into another vector to convert them to a therapeutically relevant format.
Download more information about our Protein modules:
The power of our protein modules is enhanced when used with other modules developed by Yubsis:
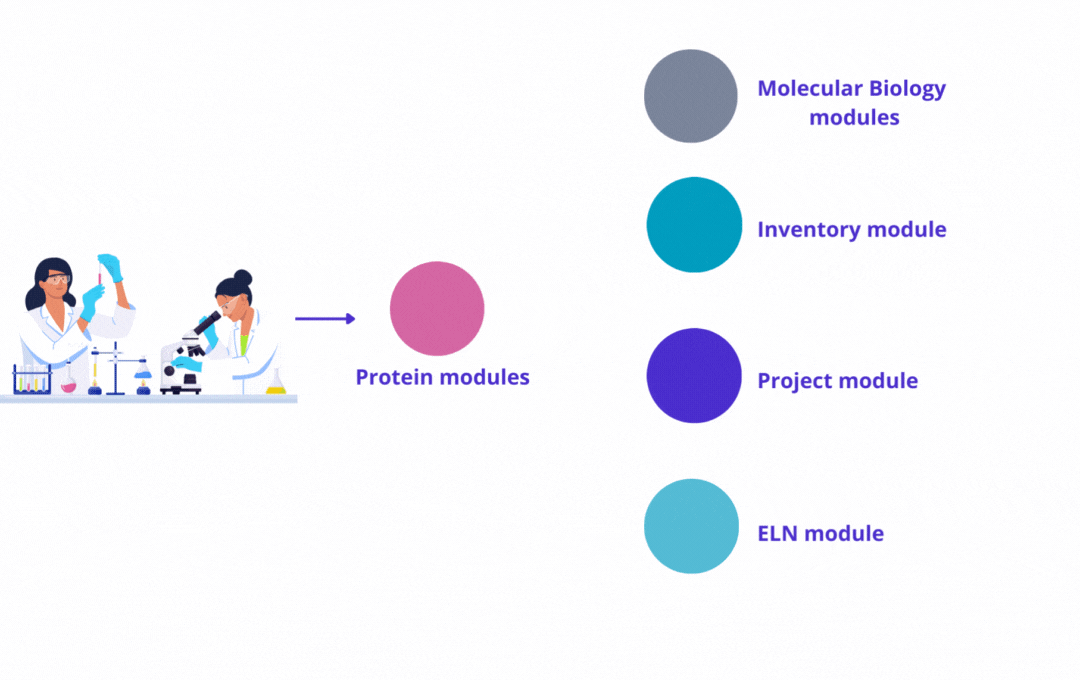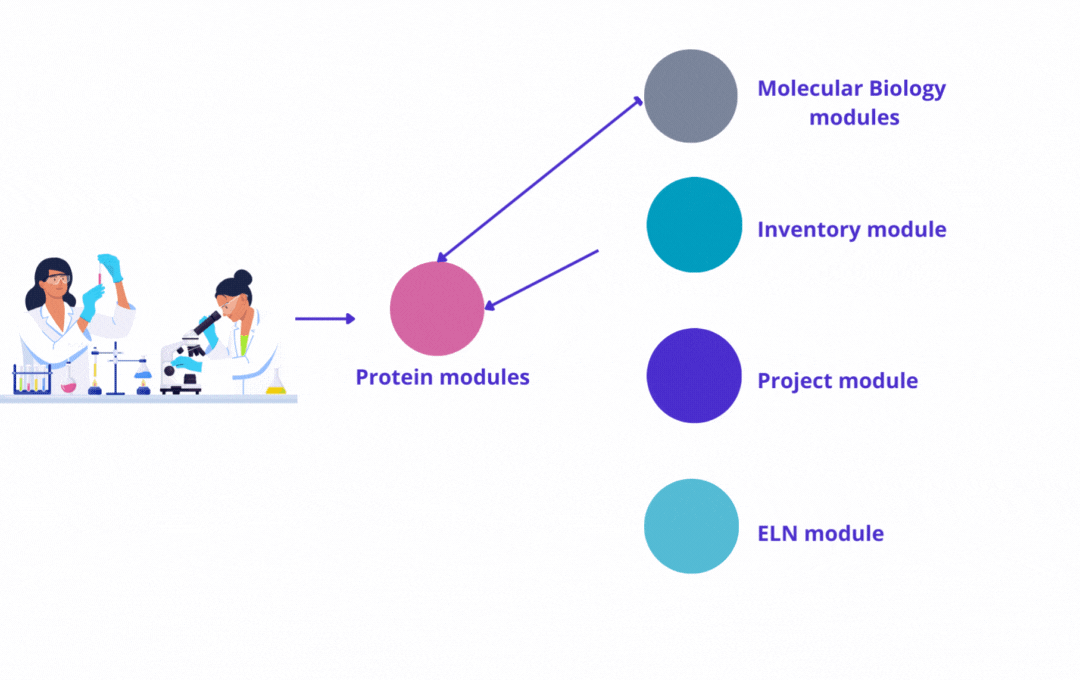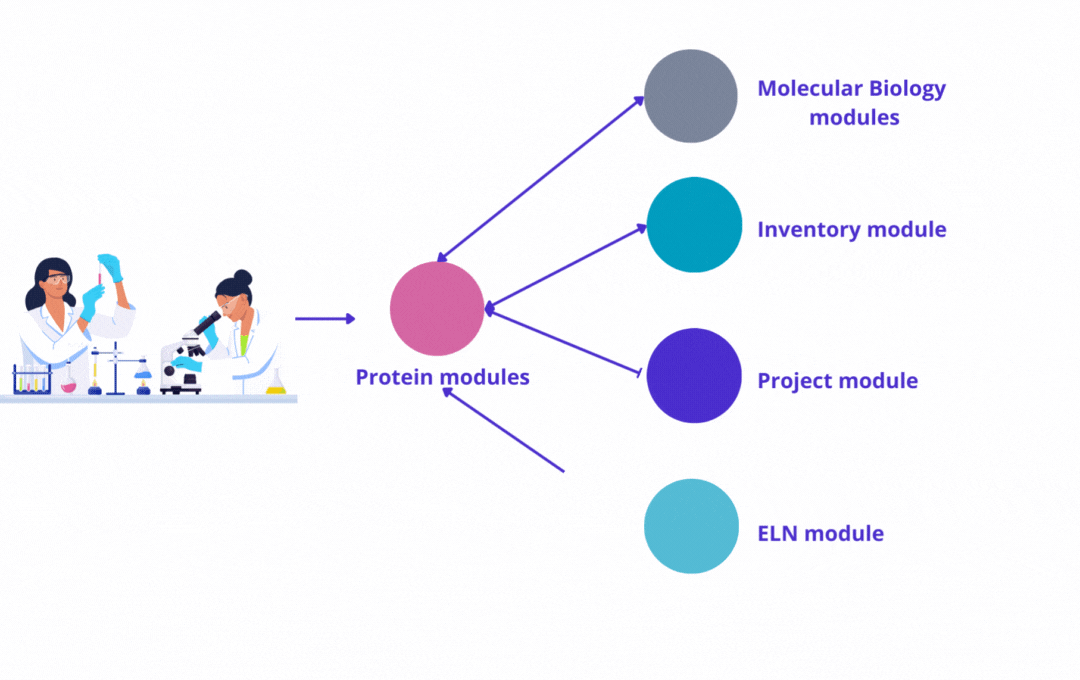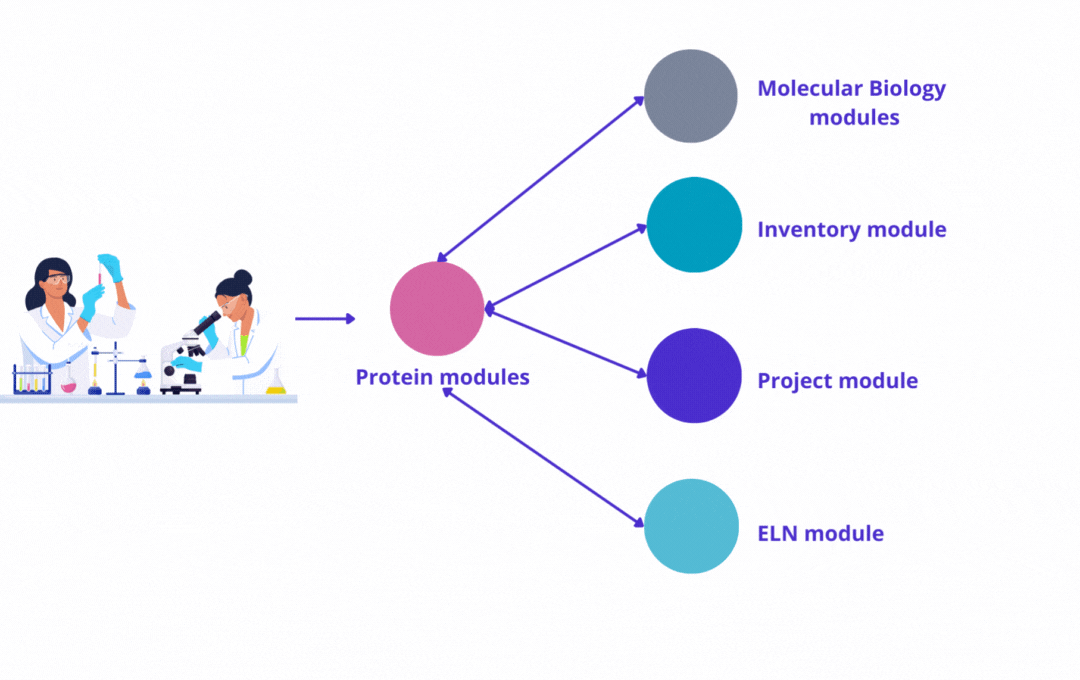
Molecular Biology
Register plasmid map, perform in silico cloning, track batches and Sanger sequencing analysis.
